DFNB1 (OMIM#: 220290)
Two different genes cause hearing loss at this locus, which maps to chromosome 13q11-q12:GJB2, encoding Connexin 26; and GJB6, encoding Connexin 30.
GJB2 (encodes Connexin 26, CX26)
CX26 (OMIM#: *121011) is a gap junction protein expressed in the supporting cells of the cochlea. The gene, GJB2, contains 2 exons, the second of which encodes the 226 amino acid protein CX26. Variants in GJB2 are found in ~50% of persons with autosomal recessive nonsyndromic hearing loss. Over 100 different variants have been identified.
GJB6 (encodes Connexin 30, CX30)
CX30 (OMIM#: *604418) is also a gap junction protein. Lerer and colleagues identified a deletion upstream of GJB2, which included the first exon of GJB6 in 2001. Del Castillo and colleagues characterized this deletion as a 342kb fragment of chromosome 13 that included D13S1830 in addition to a portion of GJB6, and so this deletion is commonly known as the del(GJB6-D13S1830) variant. It cosegregates with variants in GJB2 to cause recessively inherited deafness at the DFNB1 locus. A recent multicenter study investigating the variant spectrum in over 1500 deaf persons from 16 countries found this variant to represent 1.67% of variants at the DFNB1 locus (Van Camp, et al. 2005). A second smaller deletion, del(GJB6-D13S1854), also causes deafness at the DFNB1 locus.
Indications for screening
This test is appropriate for any patient with congenital hearing impairment with negative family history.
MORL screening
Over 97% of the identified variants at the DFNB1 locus occur in exon 2 of GJB2 (Van Camp, et al 2005). We have adopted a tiered screening process focusing first on exon 2 of GJB2 and the two GJB6-containing deletions. The finding of two deafness-causing variants is consistent with the diagnosis of hearing loss at the DFNB1 locus. If only one variant is found, variant screening for the splice site variant in exon 1 of GJB2 (IVS1 + 1 G>A) is completed. If no deafness-causing variants are found, the diagnosis of hearing loss at the DFNB1 locus is excluded based on today’s standards. (GJB2 variant screening is performed by amplification of oligonucleotide primers that flank each exon followed by bi-directional sequencing. Screening for the del(GJB6-D13S1830) and del(GJB6-D13S1854) variants is completed by PCR amplification of oligonucleotide primers flanking and within the deletion breakpoints. Products are run on agarose gel and sized to determine presence or absence of a deletion.)
Sensitivity
Sensitivity is greater than 98%.

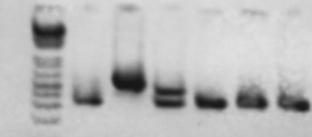
Figure legend: Lane 1, 1Kb + ladder; lane 2, wild type control; lane 3, del(GJB6-D13S1854) homozygote; lane 4, del(GJB6-D13S1830) heterozygote; lanes 5-7, normal controls.
Methodology
Sanger sequencing.
Turnaround time
Turnaround time is approximately 8 weeks (Average TAT - 29 days).
Sample required
8 - 10 cc. whole blood in lavender (EDTA) top tubes OR 10 μg DNA, minimum concentration: 50 ng/μl (A260/A280 1.8-2) resuspended in 0.1mM EDTA (10mM Tris HC1, 0.1mM EDTA, pH 8, Teknova Cat#T0220).
Cost & CPT Codes
Other Web Sites:
References
- Kelsell, D. P. et al.: Connexin 26 mutations in hereditary non-syndromic sensorineural deafness. Nature 387: 80-83, 1997. PubMed ID: 9139825
- Del Castillo, I. et al.: A deletion involving the connexin 30 gene in nonsyndromic hearing impairment. New Eng. J. Med. 346: 243-249, 2002. PubMed ID: 11807148
- Del Castillo, I. et al.: Prevalence and evolutionary origins of the del(GJB6-D13S1830) mutation in the DFNB1 locus in hearing-impaired subjects: a multicenter study. Am J Hum Genet. 2003 Dec;73(6):1452-8. Epub 2003 Oct 21. PubMed ID: 14571368
- Azaiez,H.et al.: GJB2: The spectrum of deafness-causing allele variants and their phenotype. Human Mutation 2004 Oct;24(4):305-11. PubMed ID: 15365987
- Del Castillo, F. J. et al.: A Novel Deletion Involving the Connexin-30 Gene, del(GJB6-D13S1854), Found in trans with Mutations in the GJB2 Gene (Connexin-26) in Subjects with Autosomal Recessive Non-Syndromic Hearing Impairment. J Med Genet 2005; 42: 588-594. PubMed ID: 15994881
- Snoeckx, R. L. et al.: GJB2 Mutations and Degree of Hearing Loss: A Multicenter Study. Am J Hum Genet. 77:945-957, 2005. PubMed ID: 16380907
The Clinical Diagnostics Service of the Molecular Otolaryngology and Renal Research Laboratories is a Joint Commission-approved CLIA-accredited diagnostic laboratory.